2.7.2.3.2. Content¶
| Inheritance diagram: | |
|---|---|
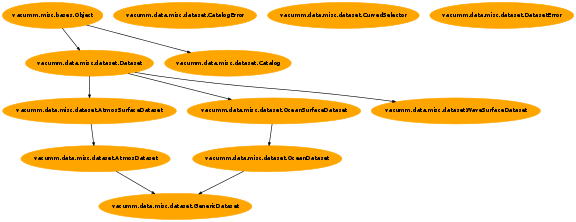
Common dataset utilities
-
class
AtmosDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.AtmosSurfaceDataset-
auto_generic_var_names= ['oro', 'wdir', 'wspd', 'uair', 'vair', 'wair', 'tair', 'pa', 'tkea']¶
-
critical= <bound class method AtmosDataset.wrapper>¶
-
debug= <bound class method AtmosDataset.wrapper>¶
-
default_altitude_search_mode= None¶
-
description= 'Generic atmospheric dataset'¶
-
domain= 'atmos'¶
-
error= <bound class method AtmosDataset.wrapper>¶
-
exception= <bound class method AtmosDataset.wrapper>¶
-
finalize_object(var, squeeze=False, order=None, asvar=None, torect=True, depthup=None, **kwargs)[source]¶ Finalize a variable
Params: - squeeze, optional: If not False, squeeze singletons axes using
squeeze_variable(). - order, optional: If not None, change the axes order of the variable. It must contains letters like ‘txyz-‘.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar_<param>: Param passed to
grow_variables(). - torect, optinal: Try to convert curvilinear grid to rectangular
grid using
curv2rect(). - depthup, optional: If not False, try the make depth positive up
using
makedepthup().
- squeeze, optional: If not False, squeeze singletons axes using
-
get_altitude(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the altitude [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates.
#not yet-
"dz": Estimate from layer thinknesses (seeget_dz()) #not yet-"axis": Read it from an axis (if not sigma coordinates)#You can specifiy a list of them:
['dz', 'sigma']#You can also negate the search with #a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_altitude_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the altitude at T location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates.
#not yet-
"dz": Estimate from layer thinknesses (seeget_dz()) #not yet-"axis": Read it from an axis (if not sigma coordinates)#You can specifiy a list of them:
['dz', 'sigma']#You can also negate the search with #a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_altitude_w(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the altitude at W location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates.
#not yet-
"dz": Estimate from layer thinknesses (seeget_dz()) #not yet-"axis": Read it from an axis (if not sigma coordinates)#You can specifiy a list of them:
['dz', 'sigma']#You can also negate the search with #a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_loglevel= <bound class method AtmosDataset.wrapper>¶
-
get_oro(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the orography for SLEVE vertical coordinates [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_pa(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the absolute pressure [Pa]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_selector(level=None, **kwargs)[source]¶ Get a cdms2.selectors.Selector from specified time/lat/lon/level selection
Params: - time/lon/lat/level, optional: Refine or set the selector with these components.
- only, optional: Work only on one component, like “time” or “t”.
- merge, optional: Merge with selector created at initialization time (stored
in attribute
selector.) - split, optional: return a splitted selector (see
split_selector())
-
get_tair(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the temperature [degrees_kelvin]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_tkea(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the turbulent Kinetic Energy [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_uair(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the zonal wind speed (westerly) [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vair(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the meridional wind speed (northerly) [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_variable(varname, level=None, squeeze=False, **kwargs)[source]¶ Load a variable in a best time serie fashion.
Params: - varname: Either a generic variable name listed in
CF_VAR_SPECS, a netcdf name with a ‘+’ a prefix, a tuple of netcdf variable names or dictionnary of specification names with a search argument ofncfind_var()(tuple of netcdf names or dictionary). - time/lon/lat/level: Selector components.
- squeeze: If true, call
squeeze_variable()on the returned variable. - order: If not None, specify the output variable axes order.
- depthup: Make depths up.
- torect: Make grid rectangular if possible.
- at/toloc: Interpolate the variable to another location on the grid
using
toloc(). Note that thearakawa_grid_typemust be defined. - format: Format the variable and its axes using
format_var()? - warn: Display a warning message if the variable can”t be retreived.
- Other kwargs are passed to
ncread_files().
Return: cdms2 variable or None if not found
Example: >>> get_variable('ssh', lon=(-10,0)) >>> get_variable('+xe') >>> get_variable(dict(search={'standard_name':'sea_surface_height_above_sea_level'}))
- varname: Either a generic variable name listed in
-
get_wair(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the upward wind speed [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_wdir(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the wind direction [degrees]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_wspd(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the wind speed [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
info= <bound class method AtmosDataset.wrapper>¶
-
is_debug= <bound class method AtmosDataset.wrapper>¶
-
is_verbose= <bound class method AtmosDataset.wrapper>¶
-
name= 'atmos'¶
-
notice= <bound class method AtmosDataset.wrapper>¶
-
notset= <bound class method AtmosDataset.wrapper>¶
-
set_loglevel= <bound class method AtmosDataset.wrapper>¶
-
verbose= <bound class method AtmosDataset.wrapper>¶
-
warning= <bound class method AtmosDataset.wrapper>¶
-
-
class
AtmosSurfaceDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.Dataset-
auto_generic_var_names= ['nethf', 'senhf', 'hfsen', 'lathf', 'hflat', 'swhf', 'lwhf', 'evap', 'rain', 'taux', 'tauy', 'u10m', 'v10m', 't2m', 'hu2m', 'z0a', 'cda', 'cha', 'cea']¶
-
critical= <bound class method AtmosSurfaceDataset.wrapper>¶
-
debug= <bound class method AtmosSurfaceDataset.wrapper>¶
-
error= <bound class method AtmosSurfaceDataset.wrapper>¶
-
exception= <bound class method AtmosSurfaceDataset.wrapper>¶
-
get_cda(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the averaged drag momentum coefficient [W s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_cea(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the averaged latent heat flux coefficient [W s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_cha(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the averaged drag thermal coefficient [W s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_evap(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the evaporation (positive when directed downward) [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_hflat(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the latent heat flux (positive when directed upward) [W.m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_hfsen(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sensible heat flux (positive when directed upward) [W m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_hu2m(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 2 m specific humidity [kg kg-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_lathf(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the latent heat flux (positive when directed downward) [W.m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_loglevel= <bound class method AtmosSurfaceDataset.wrapper>¶
-
get_lwhf(**kwargs)¶
-
get_nethf(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the net radiation [W m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_rain(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the precipitation [Downward (positive when it is raining)] [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_senhf(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sensible heat flux (positive when directed downward) [W m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_swhf(**kwargs)¶
-
get_t2m(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 2 m temperature [K]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_taux(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the surface wind stress along X [N m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_taux_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the surface wind stress along X at U location [N m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_tauy(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the surface wind stress along Y [N m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_tauy_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the surface wind stress along Y at V location [N m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u10m(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m zonal wind speed (westerly) [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u10m_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m zonal wind speed (westerly) at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u10m_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m zonal wind speed (westerly) at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u10m_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m zonal wind speed (westerly) at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v10m(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m meridional wind speed (northerly) [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v10m_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m meridional wind speed (northerly) at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v10m_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m meridional wind speed (northerly) at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v10m_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the 10-m meridional wind speed (northerly) at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_z0a(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the air roughness length [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
info= <bound class method AtmosSurfaceDataset.wrapper>¶
-
is_debug= <bound class method AtmosSurfaceDataset.wrapper>¶
-
is_verbose= <bound class method AtmosSurfaceDataset.wrapper>¶
-
name= 'atmossurface'¶
-
notice= <bound class method AtmosSurfaceDataset.wrapper>¶
-
notset= <bound class method AtmosSurfaceDataset.wrapper>¶
-
set_loglevel= <bound class method AtmosSurfaceDataset.wrapper>¶
-
verbose= <bound class method AtmosSurfaceDataset.wrapper>¶
-
warning= <bound class method AtmosSurfaceDataset.wrapper>¶
-
-
class
Catalog(files=None, filepattern=None, time=None, **kwargs)[source]¶ Bases:
vacumm.misc.bases.ObjectBuild dataset files list based on self.get_config
Variables in config (all optionnal):
- files: explicit list of datasets files
- filepattern , time: see
list_forecast_files()
These configuration can be passed when creating the Catalog object like any other Object, examples:
>>> c = Catalog(config='configfile.cfg') >>> c = Catalog(config=dict(files=('file1.nc', 'file2.nc'))) >>> c = Catalog(config=dict(filepattern='model_%Y-%m-%d', time=('2010-01-01', '2011-01-01'))) >>> c = Catalog(config=dict(filepattern='model_%Y-%m-%d', time=('2010-01-01', '2011-01-01'), files=('file1.nc', 'file2.nc')))
-
critical= <bound class method Catalog.wrapper>¶
-
debug= <bound class method Catalog.wrapper>¶
-
error= <bound class method Catalog.wrapper>¶
-
exception= <bound class method Catalog.wrapper>¶
-
classmethod
find_datasets(filepattern, time, **kwargs)[source]¶ Perform dataset search using
list_forecast_files()
-
get_datasets()[source]¶ Get the logical view of datasets.
- Datasets are specified by this object configuration (
get_config()) using: - files: use files specified directly
- filepattern and time: perform dataset search using
find_datasets()
Return: - List of strings, the dataset files found
Todo
- Ensure returned dataset order (by (run)date)
- Add a tutorial about
Catalog
- Datasets are specified by this object configuration (
-
get_loglevel= <bound class method Catalog.wrapper>¶
-
info= <bound class method Catalog.wrapper>¶
-
is_debug= <bound class method Catalog.wrapper>¶
-
is_verbose= <bound class method Catalog.wrapper>¶
-
notice= <bound class method Catalog.wrapper>¶
-
notset= <bound class method Catalog.wrapper>¶
-
set_loglevel= <bound class method Catalog.wrapper>¶
-
verbose= <bound class method Catalog.wrapper>¶
-
warning= <bound class method Catalog.wrapper>¶
-
exception
CatalogError[source]¶ Bases:
exceptions.Exception
-
class
CurvedSelector(grid, select, force=True, xid=None, yid=None)[source]¶ Bases:
objectCurved grid multiple selector
-
class
Dataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.misc.bases.ObjectGeneric dataset representation.
Handle multiple datasets (files) in a best time serie view.
Variables specifications are stored in
ncobj_specs. This global attribute must be refined for all new dataset.-
ncobj_specs¶ Dictionary describing variable specifications: final id of the variable as the keys, and aliases ad slice as the content. Here is an example:
ncobj_specs = { 'u10m':dict( search=dict(id=['u', 'u10', 'uwind', '.*u.*']), select=(Ellipsis, 0)), 'sst':dict( search=dict(standard_name="sea_surface_temperature"), select=(Ellipsis, -1)), atts=dict(units="DegC"), }
Params: - dataset: A dataset or list of dataset (see
load_dataset()). Argument to are either aCataloginstance, or compatible withlist_forecast_files(). In this latter case, please read the documentation of this function carefully. - select: A selector to be applied by default.
- time: Time for searching files (see
list_forecast_files()) which overwrites configuration. - sort/nopat/patfreq/patfmtfunc/check, optional: These arguments are passed to
list_forecast_files(). - Extra keywords are passed to Object.__init__, you may then pass a config filepath/dict/ConfigObj.
See also: Examples: Classical cases:
>>> d = Dataset("mars.*.nc") >>> d = Dataset("mars.%Y%m%d%H00.nc", time=('2000','2000-02-15')) >>> d = Dataset(['file1.nc', 'file2.nc']) >>> d = Dataset('http://opendap.net/mars.nc')
With
load_dataset():>>> d = Dataset() >>> d.load_dataset('file1.nc') >>> d.load_dataset(['file2.nc', 'file3.nc'], append=True)
Is equivalent to:
>>> d = Dataset(['file1.nc', 'file2.nc', 'file3.nc'])
Another example using configuration:
>>> d = Dataset(config='dataset.cfg')
- With configuration file ‘dataset.cfg’:
- [Dataset]
- [[Catalog]]
files = ‘file1.nc’, ‘file2.nc’, ‘file3.nc’
Note
- For a subclass of Dataset, say for example MyModel, the configuration would be:
- [MyModel]
- [[Catalog]]
files = ‘file1.nc’, ‘file2.nc’, ‘file3.nc’
Attributes: -
selector¶ A
cdms2.selectors.Selectorcreated at initialization time.
-
apply_config(config, **kwargs)[source]¶ Apply passed configuration (usually internal call from load_config)
Call
apply_config()and load datasets (if config has a Catalog section)
-
arakawa_grid_type= None¶ Arakawa grid type (see
vacumm.data.misc.arakawa)
-
auto_generic_var_names= ['corio']¶
-
critical= <bound class method Dataset.wrapper>¶
-
debug= <bound class method Dataset.wrapper>¶
-
description= None¶
-
error= <bound class method Dataset.wrapper>¶
-
exception= <bound class method Dataset.wrapper>¶
-
finalize_object(var, genname=None, format=<built-in function format>, squeeze=False, order=None, asvar=None, torect=True, atts=None, lon=None, lat=None, curvsel=None, at=None, **kwargs)[source]¶ Finalize a variable or an axis
Params: - genname, optional: Generic name to format the variable.
- format, optional: Format the variable and its axes with
format_var()? - atts, optional: Set these attributes to the variable.
- squeeze, optional: If not False, squeeze singletons axes using
squeeze_variable(). - order, optional: If not None, change the axes order of the variable. It must contains letters like ‘txyz-‘.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar_<param>: Param passed to
grow_variables(). - torect, optinal: Try to convert curvilinear grid to rectangular
grid using
curv2rect(). - lon/lat, optional: Additional spatial selection.
- curvsel, optional:
CurvedSelectorinstance. - at/toloc: Interpolate the variable to another location on the grid
using
toloc(). Note that thearakawa_grid_typemust be defined.
-
get(varname, **kwargs)[source]¶ Generic way to get a variable
It first tries to find a
get_*method, then callget_variable()if no method is found.
-
get_axis(name, select=None, select2=None, dataset=None, warn=True, getid=False, searchmode=None, format=True)[source]¶ Retreive a 1D or 2D axis.
Params: - name: Generic axis name.
- select optional: Selection along this axis. Only slices are accepted for 2D axes.
- select2 optional: Selection on the other dimension for 2D axes.
- dataset optional: find axis based on this dataset.
- getid, optional: If True, only return the id (netcdf name) or None.
- warn, optional: Display a warning in case of problem.
- searchmode, optional: Search order (see
ncfind_obj()). - format, optional: Format the axis using
format_axis()?
Return: - cdms2 axis or None if not found, or id
-
get_corio(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the coriolis parameter [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_corio_f(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the coriolis parameter at F location [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_corio_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the coriolis parameter at T location [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_corio_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the coriolis parameter at U location [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_corio_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the coriolis parameter at V location [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ctime(*args, **kwargs)[source]¶ Get time axis as a list of
cdtime.comptimeIt is a simple shortcut to:
>>> ds.get_time().asComponentTime()
Params: All arguments are passed to get_time()
-
get_dx(**kwargs)[source]¶ Get the grid resolution along X
It can be stored as an variable or computed from coordinates.
-
get_dy(**kwargs)[source]¶ Get the grid resolution along Y
It can be stored as an variable or computed from coordinates.
-
get_hsection(varname, depth, time=None, lat=None, lon=None, timeavg=False, warn=True, **kwargs)[source]¶ Get a horizontal section of a variable for a specified depth
Params: - varname: Generic var name.
- depth: Target depth.
- timeavg, optional: Time average of results.
- depth_<param>, optional: Param is passed to
get_depth(). - interp_<param>, optional: Param is passed to
interp1d().
-
get_latitude(lat=None, lon=None, **kwargs)¶ Get latitude axis
-
get_level(level=None, **kwargs)[source]¶ Get level axis, based on
get_axis()
-
get_loglevel= <bound class method Dataset.wrapper>¶
-
get_longitude(lon=None, lat=None, **kwargs)¶ Get longitude axis
-
get_resol(degrees=False, at='t', mode=None, warn=True, **kwargs)[source]¶ Get the horizontal grid resolutions
Params: - degrees, optional: In degrees or meters?
- local, optional: Get resolution at each point?
Return: dx,dySee also:
-
get_selector(time=None, lon=None, lat=None, level=None, merge=True, only=None, split=False, **kwargs)[source]¶ Get a cdms2.selectors.Selector from specified time/lat/lon/level selection
Params: - time/lon/lat/level, optional: Refine or set the selector with these components.
- only, optional: Work only on one component, like “time” or “t”.
- merge, optional: Merge with selector created at initialization time (stored
in attribute
selector.) - split, optional: return a splitted selector (see
split_selector())
-
get_seltimes(time=None)[source]¶ Get a zero to two elements time selector specifications
It is the addition of non global and local time selection specs
-
get_shape(dims='tzyx', warn=True)[source]¶ Get the dataset shape from known generic dims
Params: - dims, optional: Letters that select the generic dimensions to consider.
Return: A tuple of the size of dimensions. If a requested dim is not found, None is returned for its size.
-
get_time(time=None, var=None, ids=None, warn=True, **kwargs)[source]¶ Load time axis in a best time serie fashion.
Params: - time: time selector
-
get_time_res()[source]¶ Get the estimated time resolution, based on the two first time coordinates
Return: - resolution as datetime.timedelta
-
get_transect(varname, lons, lats, times=None, method='bilinear', subsamp=3, getcoords=False, timeavg=False, outaxis=None, time=None, lon=None, lat=None, level=None, warn=True, **kwargs)[source]¶ Get a transect between two points
It uses
transect().Params: varname: Generic var name.
lons/lats: Specification of transect, either
- Coordinates of first and last point in degrees as
tuples in the form
(lon0,lon1)and(lat0,lat1). The array of coordinates is generated usingtransect_specs(). - Or explicit array of coordinates (as scalars, lists or arrays).
- Coordinates of first and last point in degrees as
tuples in the form
times, optional: For time transect too.
subsamp, optional: Subsampling with respect to grid cell.
method, optional: Interpolation method (see
grid2xy()).getcoords, optional: Also get computed coordinates.
outaxis, optional: Output axis.
- A cdms2 axis.
Noneor'auto': Longitudes or latitudes depending on the range.'lon'or'x': Longitudes.'lat'or'y': Latitudes.'dist'or'd': Distance in km.
timeavg, optional: Time average of results.
Return: tvarortvar,tlons,tlats
-
get_variable(varname, time=None, lon=None, lat=None, level=None, atts=None, squeeze=False, order=None, asvar=None, torect=True, depthup=None, verbose=None, warn=True, searchmode='is', format=True, at=None, grid=None, **kwargs)[source]¶ Load a variable in a best time serie fashion.
Params: - varname: Either a generic variable name listed in
CF_VAR_SPECS, a netcdf name with a ‘+’ a prefix, a tuple of netcdf variable names or dictionnary of specification names with a search argument ofncfind_var()(tuple of netcdf names or dictionary). - time/lon/lat/level: Selector components.
- squeeze: If true, call
squeeze_variable()on the returned variable. - order: If not None, specify the output variable axes order.
- depthup: Make depths up.
- torect: Make grid rectangular if possible.
- at/toloc: Interpolate the variable to another location on the grid
using
toloc(). Note that thearakawa_grid_typemust be defined. - format: Format the variable and its axes using
format_var()? - warn: Display a warning message if the variable can”t be retreived.
- Other kwargs are passed to
ncread_files().
Return: cdms2 variable or None if not found
Example: >>> get_variable('ssh', lon=(-10,0)) >>> get_variable('+xe') >>> get_variable(dict(search={'standard_name':'sea_surface_height_above_sea_level'}))
- varname: Either a generic variable name listed in
-
info= <bound class method Dataset.wrapper>¶
-
is_debug= <bound class method Dataset.wrapper>¶
-
is_verbose= <bound class method Dataset.wrapper>¶
-
load_dataset(dataset, time=None, **kwargs)[source]¶ Load dataset files.
Params: - dataset: can be either:
- an instance or list of strings (filepath/filepattern/url) that
will be passed to
list_forecast_files(). - an instance of
Catalog
- an instance or list of strings (filepath/filepattern/url) that
will be passed to
- time: used with
Catalog.find_datasets()when dataset is a string (or list of strings). - Extra keywords are passed to
list_forecast_files()(viaCatalog.find_datasets()).
Keyword parameters: - append: keep previously loaded datasets if True
Return: The list of (newly) loaded datasets
Example: >>> d.load_dataset('../../data/model/mars/champs_%Y%m_BOBI.nc', ('2004-01-01', '2004-04-01'), patfreq=(2,'month'), verbose=True) [2013-01-14 09:42:43 CET Catalog INFO] Searching datasets using pattern: ['../../data/model/mars/champs_%Y%m_BOBI.nc'], time: ('2004-01-01', '2004-04-01') Guessing file list using: filepattern: ../../data/model/mars/champs_%Y%m_BOBI.nc time selector: ('2004-01-01', '2004-03-31') Found 2 files [2013-01-14 09:42:43 CET Catalog INFO] Found 2 datasets [2013-01-14 09:42:43 CET Dataset INFO] Loading datasets: [2013-01-14 09:42:43 CET Dataset INFO] - ../../data/model/mars/champs_200401_BOBI.nc [2013-01-14 09:42:43 CET Dataset INFO] - ../../data/model/mars/champs_200403_BOBI.nc
-
name= None¶
-
ncobj_specs= {}
-
notice= <bound class method Dataset.wrapper>¶
-
notset= <bound class method Dataset.wrapper>¶
-
plot_hsection(varname, depth, time=None, lat=None, lon=None, timeavg=False, title='%(long_name)s at %(depth)im', mask_land=False, **kwargs)[source]¶ Plot a horizontal section of a variable for a specified depth
Section is computed with
get_hsection().Params: - varname: Generic var name.
- depth: Target depth.
- timeavg, optional: Time average of results.
- depth_<param>, optional: Param is passed to
get_depth(). - interp_<param>, optional: Param is passed to
interp1d(). - Other arguments are passed to
map2().
-
plot_transect(varname, lons, lats, times=None, method='bilinear', timeavg=False, subsamp=3, outaxis=None, time=None, lon=None, lat=None, level=None, title='%(long_name)s along transect', minimap=None, **kwargs)[source]¶ Plot a transect between two points
Params: - varname: Generic var name.
- lons/lats/times: Specification of transect (see
get_transect()). - title, optional: Title of the figure.
- minimap, optional: If True, add a minimap showing the transect on a map; if False, display nothing; if None, display if no minimap already displayed.
- minimap_<param>, optional: Passed to
add_map_lines(). - Some params are passed to
get_transect(). - Other params are passed to the plot function
curve2()for 1D plots and,hov2()orsection2()for 2D plots.
-
set_loglevel= <bound class method Dataset.wrapper>¶
-
shape¶ Generic shape of the dataset
-
classmethod
squeeze_variable(var, spec=True)[source]¶ Squeeze a variable, preserving remaining axis
See also: vacumm.misc.misc.squeeze_variable()
-
toloc(var, loc, fromloc=None, copy=False, **kwargs)[source]¶ Interpolate a variable to another location
It has no effect if the current
Datasetinstance has no validarakawa_grid_typedefined (None).Params: - var: A CDAT array.
- loc: A physical location (see
vacumm.data.misc.arakawa.ARAKAWA_LOCATIONS). - fromloc, optional: Originating location. If
Noneit is guessed from its attributes (id, standard_name and long_name), and default toDEFAULT_LOCATION). - Extra keywords are passed to
vacumm.data.misc.arakawa.ArakawaGrid.loc2loc().
-
torect(var, curvsel=None)[source]¶ Place a variable on rectangular grid if possible using
curv2rect()Params: - var: CDAT variable or grid.
-
verbose= <bound class method Dataset.wrapper>¶
-
warning= <bound class method Dataset.wrapper>¶
-
-
exception
DatasetError[source]¶ Bases:
exceptions.Exception
-
class
GenericDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.AtmosDataset,vacumm.data.misc.dataset.OceanDatasetGeneric
Datasetclass to load everything-
critical= <bound class method GenericDataset.wrapper>¶
-
debug= <bound class method GenericDataset.wrapper>¶
-
description= 'Generic dataset'¶
-
error= <bound class method GenericDataset.wrapper>¶
-
exception= <bound class method GenericDataset.wrapper>¶
-
get_loglevel= <bound class method GenericDataset.wrapper>¶
-
info= <bound class method GenericDataset.wrapper>¶
-
is_debug= <bound class method GenericDataset.wrapper>¶
-
is_verbose= <bound class method GenericDataset.wrapper>¶
-
name= 'generic'¶
-
notice= <bound class method GenericDataset.wrapper>¶
-
notset= <bound class method GenericDataset.wrapper>¶
-
set_loglevel= <bound class method GenericDataset.wrapper>¶
-
verbose= <bound class method GenericDataset.wrapper>¶
-
warning= <bound class method GenericDataset.wrapper>¶
-
-
class
OceanDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.OceanSurfaceDataset-
auto_generic_var_names= ['temp', 'sal', 'u3d', 'v3d', 'ubt', 'vbt', 'kz', 'bathy', 'eke', 'tke']¶
-
critical= <bound class method OceanDataset.wrapper>¶
-
debug= <bound class method OceanDataset.wrapper>¶
-
default_depth_search_mode= None¶
-
description= 'Generic ocean dataset'¶
-
domain= 'ocean'¶
-
error= <bound class method OceanDataset.wrapper>¶
-
exception= <bound class method OceanDataset.wrapper>¶
-
finalize_object(var, squeeze=False, order=None, asvar=None, torect=True, depthup=None, **kwargs)[source]¶ Finalize a variable
Params: - squeeze, optional: If not False, squeeze singletons axes using
squeeze_variable(). - order, optional: If not None, change the axes order of the variable. It must contains letters like ‘txyz-‘.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar_<param>: Param passed to
grow_variables(). - torect, optinal: Try to convert curvilinear grid to rectangular
grid using
curv2rect(). - depthup, optional: If not False, try the make depth positive up
using
makedepthup().
- squeeze, optional: If not False, squeeze singletons axes using
-
get_bathy(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at T location [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at u-location
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at v-location
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_dens(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the sea water density [kg m-3]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - dens_<param>, optional: Passed to :func:`~vacumm.diag.thermdyn.density
- depth_<param>, optional: Passed to
get_depth() - mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."tempsal": Estimate from temperature and salinity.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_depth(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the depth [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates."dz": Estimate from layer thinknesses (seeget_dz())"axis": Read it from an axis (if not sigma coordinates)
You can specifiy a list of them:
['dz', 'sigma']You can also negate the search with a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_depth_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the depth at T location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates."dz": Estimate from layer thinknesses (seeget_dz())"axis": Read it from an axis (if not sigma coordinates)
You can specifiy a list of them:
['dz', 'sigma']You can also negate the search with a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_depth_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the depth at U location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates."dz": Estimate from layer thinknesses (seeget_dz())"axis": Read it from an axis (if not sigma coordinates)
You can specifiy a list of them:
['dz', 'sigma']You can also negate the search with a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_depth_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the depth at V location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates."dz": Estimate from layer thinknesses (seeget_dz())"axis": Read it from an axis (if not sigma coordinates)
You can specifiy a list of them:
['dz', 'sigma']You can also negate the search with a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_depth_w(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the depth at W location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."sigma": Estimate from sigma coordinates."dz": Estimate from layer thinknesses (seeget_dz())"axis": Read it from an axis (if not sigma coordinates)
You can specifiy a list of them:
['dz', 'sigma']You can also negate the search with a ‘-‘ sigme before:"-dz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_dz(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the ocean layer thickness [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."depth": Estimate from depth (seeget_depth())
You can also negate the search with a ‘-‘ sigme before:
"-depth".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_dz_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the ocean layer thickness at T location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."depth": Estimate from depth (seeget_depth())
You can also negate the search with a ‘-‘ sigme before:
"-depth".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_dz_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the ocean layer thickness at U location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."depth": Estimate from depth (seeget_depth())
You can also negate the search with a ‘-‘ sigme before:
"-depth".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_dz_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the ocean layer thickness at V location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."depth": Estimate from depth (seeget_depth())
You can also negate the search with a ‘-‘ sigme before:
"-depth".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_dz_w(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the ocean layer thickness at W location [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."depth": Estimate from depth (seeget_depth())
You can also negate the search with a ‘-‘ sigme before:
"-depth".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_eke(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the eddy kinetic energy [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_extrema_location(varname, xorymin, xorymax, xory, meridional=False, extrema='min', select=None)[source]¶ Get positions of min/max values of variable extremas along a straight trajectory, zonal or meridional.
Coordinates xorymin and xorymax are longitudes and xory a latitude if the section is zonal, latitudes and a longitude if the section is meridonal
Level must be defined using the select parameter.
Params: varname: variable to process
xorymin: westernmost longitude or southernmost latitude coordinate
xorymax: eastermost longitude or northernmost latitude coordinate
xory: longitude or latitude coordinate
meridional: if true, hovmoller is meridional, at a longitude and along given latitude range (default is zonal)
extrema: type of extrema, one of:
- min: retrieve minimum values positions
- max: retrieve maximum values positions
select: selector (should at least restrict to one level)
- select=dict(level=slice(-1,None),time=slice(0,2))
Return: A list containing in order:
- var(time,position): loaded variable data
- latitude(position): latitude corresponding to var’s position
- longitude(position): longitude corresponding to var’s position
Example: >>> get_extrema_location(self, 'temp', -10, -6, 47, select=dict(level=slice(-1,None))):
-
get_hovmoller(varname, xorymin, xorymax, xory, meridional=False, method='bilinear', timeavg=False, subsamp=3, outaxis=None, time=None, lon=None, lat=None, level=None, warn=True, **kwargs)[source]¶ Get a hovmoller(time,position) section data along a straight trajectory, zonal or meridional.
Warning
This method is deprecated and must be rewritten has a special case of method
get_transect().Coordinates xorymin and xorymax are longitudes and xory a latitude if the section is zonal, latitudes and a longitude if the section is meridonal
Level must be defined using the select parameter.
Params: - varname: variable to process
- xorymin: westernmost longitude or southernmost latitude coordinate
- xorymax: eastermost longitude or northernmost latitude coordinate
- xory: longitude or latitude coordinate
- meridional: if true, hovmoller is meridional, at a longitude and along given latitude range (default is zonal)
- lon/lat/level/time: Selection.
- Other keywords are passed to
get_transect().
Return: A list containing in order: - var(time,position): hovmoller variable - latitude(position): latitude corresponding to var’s position - longitude(position): longitude corresponding to var’s position
Example: >>> get_hovmoller(self, 'sst', -10, -6, 47):
-
get_ke(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the kinetic energy [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."uvgbt": Estimate from barotropic geostrophic velocity (get_uvgbt()).
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_kz(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the vertical diffusivity [m2 s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_layer(varname, depth, timeavg=True, **kwargs)[source]¶ Get an horizontal section of a variable at a specified depth
Warning
This method is now an alias for method
get_hsection()
-
get_localized_computed_values(varname, xorymin, xorymax, xory, meridional=False, operation='min', select=None)[source]¶ Get min/max/mean values of a variable along a straight trajectory, zonal or meridional.
Coordinates xorymin and xorymax are longitudes and xory a latitude if the section is zonal, latitudes and a longitude if the section is meridonal
Level must be defined using the select parameter.
Params: varname: variable to process
xorymin: westernmost longitude or southernmost latitude coordinate
xorymax: eastermost longitude or northernmost latitude coordinate
xory: longitude or latitude coordinate
meridional: if true, hovmoller is meridional, at a longitude and along given latitude range (default is zonal)
operation: type of operation, one of:
- min: retrieve minimum values
- mean: retrieve mean values
- max: retrieve maximum values
select: selector (should at least restrict to one level)
- select=dict(level=slice(-1,None),time=slice(0,2))
Return: A list containing in order: - var(time,position): loaded variable data - latitude(position): latitude corresponding to var’s position - longitude(position): longitude corresponding to var’s position
Example: >>> get_localized_computed_values(self, 'temp', -10, -6, 47, select=dict(level=slice(-1,None))):
-
get_loglevel= <bound class method OceanDataset.wrapper>¶
-
get_mixed_layer_depth(select)[source]¶ Get mixed layer depth
Warning
This method is deprecated by
get_mld().MLD is computed for each time step and then averaged
Params: - select: selector with at least a time component
Return: - mld with shape (latitude,longitude)
-
get_mld(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the mixed layer depth [m]
Params: time/level/lat/lon, optional: For selection (tuples or slices).
squeeze, optional: Squeeze singleton dimensions (see
squeeze_variable(), likeTrue,zor['x','y']).deltadens, optional: Density difference with surface
deltatemp, optional: Temperature difference with surface.
kzmax, optional: Kz max for search for low values
mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."deltatemp": Estimate from a difference in temperature."deltadens": Estimate from a difference in density."twolayers": Shalow water mode with two density layers."kz": Depth where ks becomes low.
You can specifiy a list of them:
['deltadens', 'deltatemp']You can also negate the search with a ‘-‘ sigme before:"-kz".asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables().asvar, optional: Reshape as this variable.
at, optional: Interpolate to this grid location using
Datatset.toloc().format, optional: Format the variable and its axes using
format_var().torect, optional: If possible, convert a curvilinear grid to a rectilinar grid using
curv2rect().order, optional: Change order of axes (like
'xty').Other keywords are passed to
ncread_files().
-
get_potential_energy_deficit(select)[source]¶ Get potential energy deficit
PED is computed for each time step and then averaged
Params: - select: selector with at least a time component
Return: - ped with shape (latitude,longitude)
-
get_sal(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the salinity [PSU]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_section(varname, xmin, ymin, xmax, ymax, timeavg=True, **kwargs)[source]¶ Get a (vertical) section data along a straight trajectory, not necessary zonal or meridional.
Warning
This method is deprecated by the
get_transect()method.Params: - varname: variable to process
- xmin: westernmost longitude coordinate
- ymin: southernmost latitude coordinate
- xmax: eastermost longitude coordinate
- ymax: northernmost latitude coordinate
- timeavg: if true, average date along time if needed
Return: A list containing in order: - var(level,position): section variable - depth(level) FOR 3D VARIABLES ONLY: depth corresponding to var’s level - latitude(position): latitude corresponding to var’s position - longitude(position): longitude corresponding to var’s position
-
get_selector(level=None, **kwargs)[source]¶ Get a cdms2.selectors.Selector from specified time/lat/lon/level selection
Params: - time/lon/lat/level, optional: Refine or set the selector with these components.
- only, optional: Work only on one component, like “time” or “t”.
- merge, optional: Merge with selector created at initialization time (stored
in attribute
selector.) - split, optional: return a splitted selector (see
split_selector())
-
get_ssd(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the sea surface density [PSU]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - dens_<param>, optional: Passed to :func:`~vacumm.diag.thermdyn.density
- mode, optional: Computing mode
None: Try all modes, in the following order."var": Read it from a variable."tempsal": Estimate from temperature and salinity.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_stratification_data(select, timeavg=True)[source]¶ Get stratification data
Params: - select: selector
- timeavg: if true, average date along time if needed
Return: - temp, sal, dens, depth, deltadepth with shape ([time],depth,latitude,longitude)
- densmin, densmax, densmean with shape ([time],latitude,longitude)
-
get_temp(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the temperature [degrees_celsius]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_tke(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the turbulent kinetic energy [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u3d(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along X [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u3d_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along X at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u3d_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along X at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_u3d_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along X at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ubt(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along X [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ubt_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along X at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ubt_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along X at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ubt_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along X at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ugbt(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the sea water barotropic geostrophic velocity along X [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_uvgbt(getu=True, getv=True, warn=True, mode=None, **kwargs)[source]¶ Get zonal and meridional geostrophic velocity from SSH
-
get_v3d(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along Y [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v3d_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along Y at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v3d_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along Y at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_v3d_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water velocity along Y at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_variable(varname, level=None, squeeze=False, **kwargs)[source]¶ Load a variable in a best time serie fashion.
Params: - varname: Either a generic variable name listed in
CF_VAR_SPECS, a netcdf name with a ‘+’ a prefix, a tuple of netcdf variable names or dictionnary of specification names with a search argument ofncfind_var()(tuple of netcdf names or dictionary). - time/lon/lat/level: Selector components.
- squeeze: If true, call
squeeze_variable()on the returned variable. - order: If not None, specify the output variable axes order.
- depthup: Make depths up.
- torect: Make grid rectangular if possible.
- at/toloc: Interpolate the variable to another location on the grid
using
toloc(). Note that thearakawa_grid_typemust be defined. - format: Format the variable and its axes using
format_var()? - warn: Display a warning message if the variable can”t be retreived.
- Other kwargs are passed to
ncread_files().
Return: cdms2 variable or None if not found
Example: >>> get_variable('ssh', lon=(-10,0)) >>> get_variable('+xe') >>> get_variable(dict(search={'standard_name':'sea_surface_height_above_sea_level'}))
- varname: Either a generic variable name listed in
-
get_vbt(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along Y [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vbt_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along Y at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vbt_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along Y at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vbt_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea water barotropic velocity along Y at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vgbt(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)[source]¶ Read the sea water barotropic geostrophic velocity along Y [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
info= <bound class method OceanDataset.wrapper>¶
-
is_debug= <bound class method OceanDataset.wrapper>¶
-
is_verbose= <bound class method OceanDataset.wrapper>¶
-
name= 'ocean'¶
-
notice= <bound class method OceanDataset.wrapper>¶
-
notset= <bound class method OceanDataset.wrapper>¶
-
plot_extrema_location(varname, xorymin, xorymax, xory, meridional=False, extrema='min', pmap=True, select=None, **kwargs)[source]¶ Produce 1D plot of min/max positions.
Params: Plot params: - cur_<keyword>: are passed to the section plot function
curve2()excepting those about post plotting described below - map_<keyword>: are passed to the map plot function
map2()excepting those about post plotting described below - plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
- cur_<keyword>: are passed to the section plot function
-
plot_hovmoller(varname, xorymin, xorymax, xory, meridional=False, select=None, pmap=True, **kwargs)[source]¶ Produce a hovmoller plot.
Params: see
get_hovmoller()Plot params: - hov_<keyword>: are passed to the section plot function
hov2()excepting those about post plotting described below - map_<keyword>: are passed to the map plot function
map2()excepting those about post plotting described below - plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
- hov_<keyword>: are passed to the section plot function
-
plot_layer(*args, **kwargs)[source]¶ Plot a layer
Warning
This method is deprecated by
plot_hsection().- Params:
- map_<keyword>: passed to
map2() - plot_<keyword>: passed to created map
post_plot()
- map_<keyword>: passed to
Other params are passed to
get_layer()
-
plot_localized_computed_values(varname, xorymin, xorymax, xory, meridional=False, operation='min', pmap=True, select=None, **kwargs)[source]¶ Produce 1D plot of min/max values.
Params: Plot params: - cur_<keyword>: are passed to the section plot function
curve2()excepting those about post plotting described below - map_<keyword>: are passed to the map plot function
map2()excepting those about post plotting described below - plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
- cur_<keyword>: are passed to the section plot function
-
plot_mld(select, **kwargs)[source]¶ Produce mixed layer depth map.
Params: Plot params: map_<keyword>: are passed to the map plot function
map2()exceptingthose about post plotting described below
plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
-
plot_ped(select, **kwargs)[source]¶ Produce potential energy deficit map.
Params: Plot params: - map_<keyword>: are passed to the map plot function
map2()excepting those about post plotting described below - plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
- map_<keyword>: are passed to the map plot function
-
plot_section(varname, xmin, ymin, xmax, ymax, select=None, pmap=True, **kwargs)[source]¶ Produce a section plot.
Warning
This method is deprecated by
plot_transect().Params: see
get_section()Plot params: - sec_<keyword>: are passed to the section plot function
section2()excepting those about post plotting described below - map_<keyword>: are passed to the map plot function
map2()excepting those about post plotting described below - plot_[show|close|savefig|savefigs]: are passed to the post plotting function
post_plot()at end of plotting operations
Todo
- add lat/lon position lines indicator
- sec_<keyword>: are passed to the section plot function
-
plot_trajectory_map(lon, lat, **kwargs)[source]¶ Plot the “legend” map of a trajectory using
add_map_lines()Params: - lon/lat: Coordinates (in degrees) as 1D arrays.
Todo
- replace this method usage by vacumm.misc.plot.add_map_lines
-
set_loglevel= <bound class method OceanDataset.wrapper>¶
-
verbose= <bound class method OceanDataset.wrapper>¶
-
warning= <bound class method OceanDataset.wrapper>¶
-
-
class
OceanSurfaceDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.Dataset-
auto_generic_var_names= ['sst', 'sss', 'ssh', 'usurf', 'vsurf']¶
-
critical= <bound class method OceanSurfaceDataset.wrapper>¶
-
debug= <bound class method OceanSurfaceDataset.wrapper>¶
-
error= <bound class method OceanSurfaceDataset.wrapper>¶
-
exception= <bound class method OceanSurfaceDataset.wrapper>¶
-
get_loglevel= <bound class method OceanSurfaceDataset.wrapper>¶
-
get_ssh(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface height [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_sss(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface salinity [PSU]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_sst(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface temperature [degrees_celsius]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_usurf(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along X [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_usurf_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along X at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_usurf_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along X at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_usurf_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along X at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vsurf(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along Y [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vsurf_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along Y at T location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vsurf_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along Y at U location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vsurf_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface velocity along Y at V location [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
info= <bound class method OceanSurfaceDataset.wrapper>¶
-
is_debug= <bound class method OceanSurfaceDataset.wrapper>¶
-
is_verbose= <bound class method OceanSurfaceDataset.wrapper>¶
-
name= 'oceansurface'¶
-
notice= <bound class method OceanSurfaceDataset.wrapper>¶
-
notset= <bound class method OceanSurfaceDataset.wrapper>¶
-
set_loglevel= <bound class method OceanSurfaceDataset.wrapper>¶
-
verbose= <bound class method OceanSurfaceDataset.wrapper>¶
-
warning= <bound class method OceanSurfaceDataset.wrapper>¶
-
-
class
WaveSurfaceDataset(dataset=None, time=None, lon=None, lat=None, level=None, ncobj_specs=None, nopat=False, patfreq=None, patfmtfunc=None, sort=True, check=True, **kwargs)[source]¶ Bases:
vacumm.data.misc.dataset.Dataset-
auto_generic_var_names= ['hs', 'mss', 'mssx', 'mssy', 'mss', 'dir', 'fp', 't0m1', 'lm', 'ubot', 'uubr', 'vubr', 'bhd', 'foc', 'utwo', 'vtwo', 'utaw', 'vtaw', 'uuss', 'vuss', 'utus', 'vtus', 'fbb', 'utbb', 'vtbb', 'mapsta', 'bathy', 'wlv', 'ucur', 'vcur', 'uwnd', 'vwnd', 'dp', 'cha', 'utaw', 'vtaw']¶
-
critical= <bound class method WaveSurfaceDataset.wrapper>¶
-
debug= <bound class method WaveSurfaceDataset.wrapper>¶
-
error= <bound class method WaveSurfaceDataset.wrapper>¶
-
exception= <bound class method WaveSurfaceDataset.wrapper>¶
-
get_bathy(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_t(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at T location [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_u(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at u-location
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bathy_v(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the bathymetry at v-location
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_bhd(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the radiation pressure (Bernoulli head) [N s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_cha(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the averaged drag thermal coefficient [W s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_dir(**kwargs)¶
-
get_dp(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the peak direction [degrees]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_fbb(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the wave dissipation in bottom boundary layer [W m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_foc(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the wave mixing kinetic turbulent energy due to surface breaking wave [W m-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_fp(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the peak frequency [s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_hs(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the significant wave height [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_lm(**kwargs)¶
-
get_loglevel= <bound class method WaveSurfaceDataset.wrapper>¶
-
get_mapsta(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the status map [1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_mss(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the mean square slope [m m-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_mssx(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the eastward mean square slope [m m-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_mssy(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the northward mean square slope [m m-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_t0m1(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the mean wave period [s]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_ubot(**kwargs)¶
-
get_ucur(**kwargs)¶
-
get_utaw(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the eastward wave supported wind stress [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_utbb(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the zonal component of the bottom wave-ocean momentum flux [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_utus(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the eastward stokes transport [m2 s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_utwo(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the zonal component of the surface wave-ocean momentum flux [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_uubr(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the rms of near bottom zonal velocities [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_uuss(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the eastward surface stokes drift [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_uwnd(**kwargs)¶
-
get_vcur(**kwargs)¶
-
get_vtaw(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the northward wave supported wind stress [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vtbb(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the meridional component of the bottom wave-ocean momentum flux [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vtus(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the northward_stokes_transport [m2 s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vtwo(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the meridional component of the surface wave-ocean momentum flux [m2 s-2]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vubr(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the rms of near bottom meridional velocities [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vuss(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the northward surface stokes drift [m s-1]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
get_vwnd(**kwargs)¶
-
get_wlv(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)¶ Read the sea surface height above sea level [m]
Params: - time/level/lat/lon, optional: For selection (tuples or slices).
- squeeze, optional: Squeeze singleton dimensions
(see
squeeze_variable(), likeTrue,zor['x','y']). - mode, optional: Retreiving mode
"var": Get from netcdf variable only."stag": Get it from another grid location.
- asvar, optional: Grow variable to match the
asvarvariable, usinggrow_variables(). - asvar, optional: Reshape as this variable.
- at, optional: Interpolate to this grid location using
Datatset.toloc(). - format, optional: Format the variable and its axes using
format_var(). - torect, optional: If possible, convert a curvilinear grid to
a rectilinar grid using
curv2rect(). - order, optional: Change order of axes (like
'xty'). - Other keywords are passed to
ncread_files().
-
info= <bound class method WaveSurfaceDataset.wrapper>¶
-
is_debug= <bound class method WaveSurfaceDataset.wrapper>¶
-
is_verbose= <bound class method WaveSurfaceDataset.wrapper>¶
-
name= 'wave'¶
-
notice= <bound class method WaveSurfaceDataset.wrapper>¶
-
notset= <bound class method WaveSurfaceDataset.wrapper>¶
-
set_loglevel= <bound class method WaveSurfaceDataset.wrapper>¶
-
verbose= <bound class method WaveSurfaceDataset.wrapper>¶
-
warning= <bound class method WaveSurfaceDataset.wrapper>¶
-
-
check_mode(validmodes, modes, strict=False)[source]¶ Check at least one of the checked modes is valid
Params: - validmodes: Valid modes as a single or list of strings.
- modes: Modes to check.
Noneis accepted. - strict, optional: If a checked mode is
None, it returnsTrueif strict isTrueelseFalse.
Example: >>> check_mode('var', 'var') True >>> validmodes = ['var', 'depth'] >>> check_mode(validmodes, 'toto') False >>> check_mode(validmodes, None) True >>> check_mode(validmodes, 'var') True >>> check_mode(validmodes, 'var', 'toto') True >>> check_mode(validmodes, ['var', 'toto']) True >>> check_mode(validmodes, '-axis') True
-
cls¶
-
getvar_decmets(cls)[source]¶ Generate methods such as
get_sst()based on theauto_generic_var_namesattributes that provides a list of generic names of variablesThese generic names must be the keys of
vacumm.data.cf.GENERIC_VAR_SPECS. The docstring of the each method is formatted usinggetvar_fmtdoc().Methods that already exist are not overwritten.
When a generic name does not start with a “+”, its name appended with a location suffix (‘_t’, ‘_u’, ‘_v’, ‘_w’, ‘_f’) is also treated.
-
getvar_fmtdoc(func, **kwargs)[source]¶ Format the docstring of methods that get variables
It uses the
getvar_fmtdoc_tpltemplate.Params: - func: Method (or function) name that must be in the form
get_<name>, where<name>must be a generic var names invacumm.data.cf.GENERIC_VAR_SPECS. - Extra keywords define extra options in the docstring. Keys must correspond to a keyword name, and values correspond to a keyword description.
Example: # Manual way getvar_fmtdoc(OceanDataset.get_sst, extra_param='Its description') # Decorator way @getvar_fmtdoc def get_sst(self, *args, **kwargs): ...
- func: Method (or function) name that must be in the form
-
getvar_fmtdoc_tpl= "%(func_name)s(time=None, level=None, lat=None, lon=None, squeeze=False, order=None, asvar=None, at=None, format=None, torect=True, verbose=None, warn=None, mode=None, **kwargs)\n\n Read the %(long_name)s %(units)s\n\n :Params:\n\n - **time/level/lat/lon**, optional: For selection (tuples or slices).\n - **squeeze**, optional: Squeeze singleton dimensions\n (see :func:`~vacumm.misc.misc.squeeze_variable`, like\n ``True``, ``z`` or ``['x','y']``).\n %(other_params)s- **asvar**, optional: Grow variable to match the ``asvar`` variable,\n using :func:`~vacumm.misc.misc.grow_variables`.\n - **asvar**, optional: Reshape as this variable.\n - **at**, optional: Interpolate to this grid location using\n :meth:`Datatset.toloc`.\n - **format**, optional: Format the variable and its axes using\n :func:`~vacumm.data.cf.format_var`.\n - **torect**, optional: If possible, convert a curvilinear grid to\n a rectilinar grid using :func:`~vacumm.misc.grid.regridding.curv2rect`.\n - **order**, optional: Change order of axes (like ``'xty'``).\n %(kwargs_params)s\n "¶ Template for auto formatting methods that get variables (like
get_sst())
